LINKED PAPER
Phylogeography and diversification of the Dead Sea Sparrow (Passer moabiticus) in Iran: insights from a multilocus approach. Shams, B., Pons, J. M., Abdelkrim, J., & Fuchs, J. 2021. IBIS. DOI: 10.1111/ibi.12957. VIEW
Charles Darwin once wrote that ‘No one definition has yet satisfied all naturalists; yet every naturalist knows vaguely what he means when he speaks of a species.’ These words are still relevant today as taxonomists continue to argue about drawing species boundaries across the continuous variation in nature (Donald 2021). Depending on the preferred species concept, different researchers might come to opposite conclusions. For example, some birds might look morphologically distinct even though they are genetically indistinguishable. In recent years, however, taxonomy has become more pluralistic through the application of several species concepts (Ottenburghs 2019). By combining numerous lines of evidence, such as morphology, genetics and behaviour, ornithologists can make better informed decisions on the classification of birds (Padial et al. 2010). Recently, Bita Shams and her colleagues applied this integrative approach to the Dead Sea Sparrow (Passer moabiticus). Are we dealing with one species or two?
Two subspecies
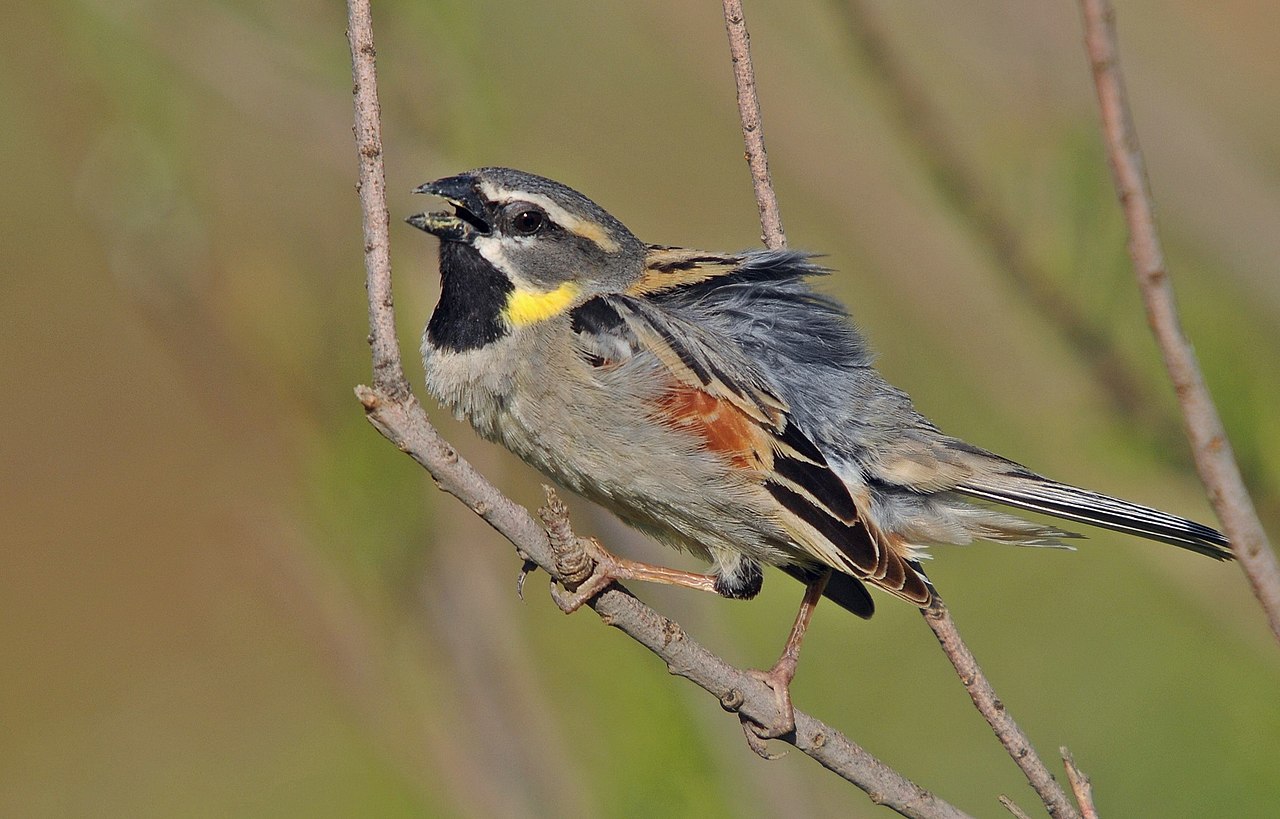
Currently, the Dead Sea Sparrow contains two subspecies that differ in distribution and plumage patterns. The subspecies P. m. moabiticus can be found from Turkey to western Iran, whereas the other subspecies P. m. yatii (also known as the Afghan Sparrow) occurs at the border between Iran and Afghanistan. Overall, the Afghan Sparrow has a lighter plumage palette and sports more extensive yellowish spots on the sides of its throat (Kirwan 2004). The researchers collected DNA samples for 51 individuals to investigate whether these differences are also reflected on the genetic level.
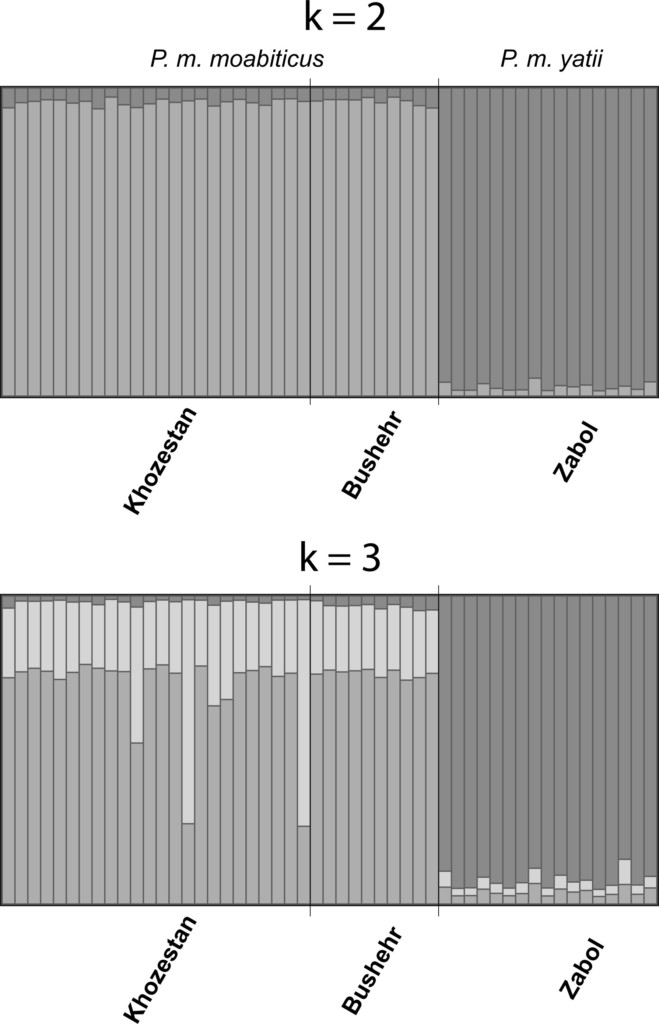
Figure 1. Genetic analyses pointed to two clear clusters (top graph) that correspond to the subspecies P. m. moabiticus (light gray) and P. m. yatii (dark gray). Extending the analyses to three clusters (bottom graph) did not reveal any population structure.
Arid conditions
Analyses of the mitochondrial gene ND2 revealed two distinct clusters corresponding to the two subspecies. A set of seven microsatellites corroborated this pattern and indicated that the subspecies have been isolated for at least 300,000 years. During this period of isolation – probably instigated by a period of extreme aridity (Abed & Yaghan 2000) – there has been little to no gene flow across the 1200 kilometre-wide gap that currently separates the populations of P. m. moabiticus and P. m. yatii. The genetic results nicely complement the other lines of evidence, prompting the researchers to propose a species rank for both subspecies: ‘moabiticus and yatii can be considered two distinct species because of diagnostic differences in plumage coloration, distinctive geographical distribution and significant genetic divergence in both mitochondrial and nuclear DNA.’ Hence, birdwatchers might have a new species to tick on their list, but the birds themselves won’t notice the difference.
References
Abed, A.M. & Yaghan, R. (2000). On the paleoclimate of Jordan during the last glacial maximum. Palaeogeography, Palaeoclimatology, Palaeoecology. 160: 23– 33. VIEW
Donald, P. F. (2021). Accounting for clinal variation and covariation in the assessment of taxonomic limits: why we should remember the ‘rules’. Ibis 163: 1106-1109. VIEW
Kirwan, G.M. (2004). The taxonomic position of the Afghan Scrub Sparrow Passer (moabiticus) yatii. Sandgrouse 26: 105– 111. VIEW
Ottenburghs, J. (2019). Avian species concepts in the light of genomics. In: Avian genomics in ecology and evolution (pp. 211-235). Springer, Cham. VIEW
Padial, J. M., Miralles, A., De la Riva, I., & Vences, M. (2010). The integrative future of taxonomy. Frontiers in Zoology, 7(1), 1-14. VIEW
Image credits
Top right: Dead Sea Sparrow (Passer moabiticus) | Dûrzan Cîrano | CC BY-SA 4.0 Wikimedia Commons
Blog posts express the views of the individual author(s) and not those of the BOU.
If you want to write about your research in #theBOUblog, then please see here




